Science cluster & challenges

Summary
The project Implementing FAIRness in structure-based drug design through Fragalysis Cloud aims to enhance the transparency and accessibility of structure-based drug design (SBDD) data by making it fully open and compliant with FAIR principles. Leveraging Diamond’s XChem service and the Fragalysis Cloud platform, the project will streamline data deposition and establish a first-of-its-kind tool for sharing, exploring, and evolving SBDD experiments. This will significantly lower the barrier to open data sharing, advancing drug discovery methodologies.
Challenge
Open Science project, Open Science Service, Cross-domain/Cross-RI
There is a strong consensus in the field of medicinal chemistry, that existing datasets for exploring and improving structure-based drug design (SBDD) are highly inadequate. The state-of-the-art of data availability has evolved into the separate repositories of the Protein Data Bank (accessed through PDBe in ELIXIR), and the ChEMBL database. These are individually well-developed and highly FAIR, but only for very specialised informaticians, and with necessarily strictly constrained ontologies. Moreover, data is often fragmented across various tools and hidden in unstructured formats, making it difficult for researchers to access and build upon existing datasets
Solution
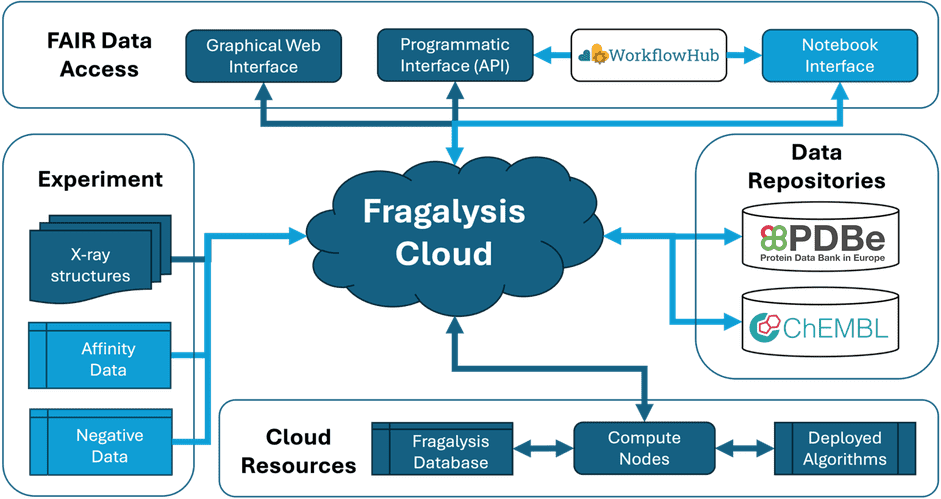
The project aims to enhance the Fragalysis Cloud platform to facilitate open and FAIR-compliant sharing of SBDD data. Fragalysis Cloud is a collaboration platform developed at Diamond’s XChem facility, used for curating, sharing and disseminating views of 3D data, and implementing best-practice medicinal chemistry algorithms and workflows for progressing results from fragment screens through early DMTA cycles. The project will map out clearly how Fragalysis can be a platform for collectively annotating and depositing data to PDBe and ChEMBL. Furthermore, it will make details of annotation workflows and computations accessible from those repositories, and generalised workflows will be made available to WorkflowHub.
By integrating data from 50–150 experiments, the project will create an accessible mechanism for depositing and querying large-scale datasets. Collaborating with key INSTRUCT (XChem) and ELIXIR services (PDBe and ChEMBL), which host the key data categories of the field (3D structures of protein-compound complexes and assay results of the compounds, including biochemical activity, biophysical affinity and ADME measurements), the project will enable researchers to explore SBDD projects with comprehensive data on context, provenance, design rationales and inspection of computational analyses.
Scientific Impact
The project will revolutionise the accessibility and reuse of SBDD data, providing an innovative tool for examining collective datasets in a FAIR manner. By making previously isolated data (and metadata) available for querying across projects, the project will improve transparency and reproducibility of experiments, while allowing to advance methodology and algorithms in the field. Researchers and developers will benefit from the insights gained from shared workflows, while contributing significantly to the FAIR data landscape in the life sciences.
The scientific relevance of the tools will be stress-tested by applying the new tools to real-world XChem data, both historic and new, and training the user community in their use, thereby greatly expanding the quantity and quality of SBDD data in the public domain.
Open science added value
Results
Three key innovations were delivered.
- XChemAlign: software that automatically transforms complex crystallographic data into biologically meaningful formats, enabling researchers to directly compare how different molecules bind to their target proteins to fully exploit potential discovery avenues.
- A data sharing infrastructure: comprehensive API endpoints and Python tools that provide programmatic access to structural and activity data, enabling automated analysis, seamless integration with computational workflows, and streamlined data preparation for deposition to PDBe.
- An interactive analysis environment: the integration of ready-to-use computational tools — Fragmenstein, HIPPO, and Syndirella — directly within the platform through Jupyter notebooks, eliminating the technical barriers that typically prevent medicinal chemists from using advanced drug design algorithms.
The project successfully processed and disseminated data for nine viral protein targets through the AI-driven Structure-enabled Antiviral Platform - ASAP consortium for Open Science antiviral drug discovery, supporting Open Science efforts against COVID-19, Zika, Dengue, Enterovirus, Chikungunya, and other emerging viral threats. A collaboration with PDBe was established to standardise fragment screening data deposition workflows, now formalised in the OpenBind partnership.
The Fragalysis Cloud platform is accessible at fragalysis.diamond.ac.uk
Publications
- Ni, X., Richardson, R.B., Godoy, A.S. et al. Combined crystallographic fragment screening and deep mutational scanning enable discovery of Zika virus NS2B-NS3 protease inhibitors. Nat Commun 16, 8930 (2025). DOI: https://doi.org/10.1038/s41467-025-63602-z
- Preprint articles on fragment screens of Coxsackievirus A16, Enterovirus D68, and Zika virus.
Final video
Principal investigator

Warren Thompson is a Senior Computational Scientist at XChem, Diamond Light Source that works on integrating software engineering, chemistry, and prototyping to make open-source fragment-progression technologies, that include structural data dissemination, FAIR-data, collaboration tools and digitised chemistry, accessible to the fragment progression community.